Table of Contents
Data preparation, cleaning, and feature encoding
All observations with a transplant date earlier than November 8, 2022 were included. All waitlist (THORACIC_WL_DATA), peri-transplant (THORACIC_DATA), and follow-up features (THORACIC_FOLLOWUP_DATA) with at least 1 observation were initially considered, yielding 786 features from the STAR File Data Dictionary. All features were reviewed for outliers and unexpected values with consultation from a committee of transplant pulmonologists. Outlier data were treated as missing. Binary features were numerically binarized as 0 or 1. Categorical features were ordinalized if they represented a scale or one-hot encoded. Waitlist and follow-up features were collected at various points in time for each patient. For waitlist data only the first and most recent values were collected. Follow-up observations closest to and within ± 3 months of one-, three-, and five-years post-transplant were used.
Principal components analysis
The features and resultant dataframe obtained from Data Preparation and Feature Encoding were further filtered for those that were missing < 10% of data, resulting in 363 features. Numerical features were mean imputed while categorical features were mode imputed. With this dataframe, principal components analysis was applied using scikit-learn29 in Python, with the top 50 principal components chosen for initial exploration. Matplotlib30 and seaborn31 were used for data exploration and visualization.
Statistical tests
To determine whether continuous variables change with respect to time (transplant date), logistic regression was performed with statsmodels32 to determine R-squared coefficients and p-values. To analyze differences in categorical variables between groups, the chi-squared test (scipy.stats)33 was used with a Benjamini–Hochberg adjustment for multiple comparisons (FDR < 0.05) (statsmodels)32. To analyze differences in continuous variables between groups, the wilcoxon rank sum test (scipy.stats)33 was used with a Benjamini–Hochberg adjustment for multiple comparisons (FDR < 0.05) (statsmodels)32. To draw statistical comparisons between the bootstrapped AUROCs between models (50 bootstraps), a Student’s t-test was performed after assessing normality with the Shapiro–Wilk test (scipy.stats)33. A Benjamini–Hochberg adjustment was made for multiple comparisons (FDR < 0.05) (statsmodels)32.
Data preparation for modeling
Feature selection and preparation for modeling
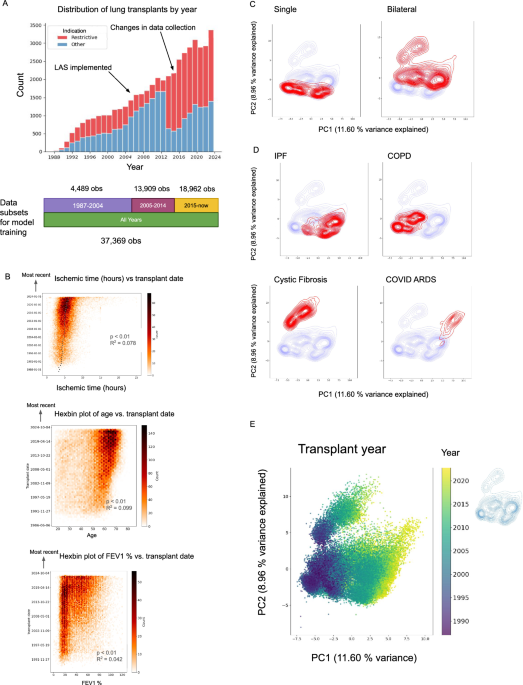
Subsets of data were chosen for modeling based on observations from the following time frames (inclusive): 1987–2004, 2005–2014, and 2015-Present. These time-related subsets were chosen due to changes in data collection illustrated in S1. All waitlist and peri-transplant data were included for all modeling tasks. Features were filtered for those that were missing < 10% of observations within each time period, resulting in 140, 145, 152 and 140 features for data from 1987–2004, 2005–2014, 2015-Present, and all years. Numerical features were mean imputed while categorical features were mode imputed.
EHRFormer feature representation and pretraining
From each of the 4 dataframes generated as described in the previous paragraph, 4 different models were pretrained from a tabular BERT initialized with random weights, available via Huggingface in the transformers package34. For the base architecture of this model (EHRFormer), we set the standard Transformer Encoder with three transformer layers, with a layer size of 64, an intermediate layer size of 128, and 4 attention heads. For input representation of the tabular data, all numerical values were quantile-binned to represent data from a single patient. Binary features were set to either the lowest or highest quartile bin. Following the standard BERT procedure, we added a 64-dimensional learnable CLS embedding at the beginning of the sequence. An additional bin was included to represent the CLS token for each patient as well as an additional bin for missing data. Each bin and each EHR feature was then assigned with a learnable 64-dimensional vector embedding. We later represented a single EHR entry with a sequence of the length of the number of features, where each element of a sequence was a 64-dimensional vector obtained by summing the feature embedding vector and its assigned bin embedding vector for every feature in the training data. In line with the BERT pretraining procedure, we pre-trained EHRFormer using a masked language model objective. The observations were divided into an 80/20 train/test split for pre-training. During this phase, we randomly masked 15% of the values in the 80% train of each entry by replacing the true bin embedding with a learnable MASK embedding. We later equipped EHRFormer with the task of predicting the actual bin values of the masked entries based on the unmasked EHR feature values. None of the observations in the 20% holdout test set were seen during pretraining.
Modeling one-, three-, and five-year mortality and lung function outcomes
Mortality label retrieval
To obtain the correct labels for one-, three-, and five-year mortality, we used the patient’s date of death (COMPOSITE_DEATH_DATE) to confirm patient death. We then calculated survival in days by subtracting the difference between the patient’s date of death and date of transplant (TX_DATE). Survival time was then used to further identify which patients had died within the one-, three-, and five-year outcomes of interest. To retrieve patients who were alive at one, three, and five years, we used the “patient status”, patient status date, and date of death variables (PX_STAT = “A “ for alive, PX_STAT_DATE, COMPOSITE_DEATH_DATE = NA). PX_STAT_DATE was used to determine total known survival time. We then added this subset of patients to patients who might have died but whose survival exceeded the 1-, 3-, and 5-year outcomes of interest for each task. To further ensure the model was not relying on shortcut features, we removed patients who died during or within 90 days of their index stay. This was determined by filtering out patients whose length of stay (LOS) exceeded survival time by at least 90 days.
Lung function < 70% of predicted label retrieval
Lung function thresholds of 70–80% have been widely adopted from general population studies of obstructive lung disease35,36. Transplant-specific factors, including laterality and transplant type, may justify lower thresholds in some contexts; single lung transplant recipients typically achieve lower peak FEV1 values (approximately 50%) compared to bilateral recipients (approximately 70%).37,38,39 Given that our cohort includes predominantly bilateral transplant recipients, we selected the 70% threshold as a compromise to leverage established risk stratification from the broader respiratory literature. To obtain labels for lung function < 70% of predicted, we used the follow-up feature FEV_percent available at 1 year post transplant (+ /- 3 months). For patients in which FEV_percent at 1 year post transplant was unavailable, we used the spiref40 package with GLI-201241 reference values to determine a calculated FEV_percent of predicted based on their absolute FEV1(L) at one year (FEV). Downsampling of the majority class was also applied as described in detail in the subsequent section EHRFormer fine-tuning and evaluation.
EHRFormer fine-tuning and evaluation
We used fine-tuning of the pretrained EHRFormer for binary classification of the patient’s mortality and lung function at 1 year. Specific outcome retrieval is described above. To perform this task within specific time frames, we used one of the 4 pretrained models that corresponded to these time frames: 1987–2004, 2005–2014, 2015-Present, and all years. With the data and pretrained model from data spanning all years, we also performed identical binary classification of whether the patient was alive or dead at 3 and 5 years. Due to class imbalance for all tasks, the majority class was downsampled at random to result in a 1:1 negative to positive class ratio in both the training and test sets separately.
For each task, following downsampling, hyperparameter tuning was performed using optuna’s42 Parzen Tree based estimator (objective = accuracy, direction = maximize, n_trials = 75), on the 80% train split from pretraining. We searched for optimal hyperparameters within the following space: learning_rate = [1e-6, 1e-2], per_device_train_batch_size = [16, 32, 64, 128, 256], and weight_decay = [1e-4, 1e-1]. With the tuned hyperparameters, we performed fivefold cross-validation on the 80% train split from pretraining. Splits and performance metrics (accuracy, AUROC, F1, precision, recall, and specificity) were determined using scikit-learn29. The tuned models were finally evaluated on their performance using the remaining 20% holdout test set that were not seen during pre-training, hyperparameter tuning, or fivefold cross-validation. To determine the variability of performance metrics within the test set, 50 random bootstraps were performed on the test set when calculating model metrics.
XGBoost evaluation
We similarly used XGBoost for binary classification of the patient’s mortality and lung function at 1 year within time frames corresponding to 1987–2004, 2005–2014, 2015-Present, and all years. We also performed identical binary classification of whether the patient was alive or dead at 3 and 5 years. Additional tasks included assessing the performance of XGBoost on all features, features only available at or before the time of transplant, and on the LAS. For these additional tasks, center-specific features were also included (ie. CTR_CODE, see Table S1). Where center code features were included, the Catboost Encoder package was used to perform target encoding of center codes. Finally, XGBoost was also used to determine prediction performance on single and bilateral lung transplant recipients separately. Downsampling of the majority class was done as was done for EHRFormer. Since there is no pre-training process, we divided entire datasets into an 80/20 train/test split at random. fivefold cross-validation was performed within the training set with scikit-learn. Hyperparameter tuning was performed using a Bayesian optimizer with a Gaussian Process based estimator (scikit-optimize’s BayesSearchCV)43. The search space was defined as follows: learning_rate = (0.01, 1.0), max_depth = (2, 12), subsample = (0.1, 1.0), colsample_bytree = (0.1, 1.0), reg_lambda = (1e-9, 100), reg_alpha = (1e-9, 100), min_child_weight = (1, 10), gamma = (0, 5), n_estimators = (50, 1000). To determine the variability of performance metrics within the test set, 50 random bootstraps were performed on the test set when calculating model metrics. For interpretability and insight into XGBoost model decisions, we used SHAP (SHapley Additive exPlanations)44.
EHRFormer perturbations
To gain model insights from EHRFormer, we developed a pretrained tabular BERT similar to the pretraining process described in the modeling tasks. We specifically included data from all years and included additional features of interest such as transplant year (TX_YEAR) as well as all the features from the 2015-Present model. The entire dataset was used for pretraining as opposed to setting aside a test cohort. Hyperparameter search was performed within the entire dataset as described previously. We then ran this new model on the fine-tuning binary classification task of 1-year mortality. To understand feature importance, we randomly sampled half of the input observations 10 times. From these sampled observations, we manipulated features of interest by manually changing the bins for one or multiple features in each of the sampled observations. These new “perturbed” inputs were fed to the model during fine-tuning. This process returned new probabilities of a positive class outcome (ie. 1 year mortality) determined by the model when given a perturbed set of features, visualized here as probability distributions before and after the perturbation was applied.
Hierarchical clustering for length of stay analysis
Hierarchical clustering was performed on all observations using all the features that were used for prediction of 1 year mortality (140 features). Numerical features were mean imputed while categorical features were mode imputed. The ward linkage method and euclidean distance were used. Whether an individual died during their index stay was used as the row annotation feature.
Propensity score matching for creating matched cohorts for length of stay analysis
To examine associations between those who died during the index stay and those who did not, we performed propensity score matching with psmpy45 to generate a matched cohort of controls. 1:5 matching was performed based on patients with similar transplant year, indication grouping, gender, age, and ethnicity. Missing data were imputed prior to matching with a simple mean strategy. KNN matching with propensity logits was performed at a 1:5 case:control ratio to mitigate class imbalance.